Common DFOS tools:
Documentation
|
| dfos
= Data Flow Operations System, the common tool set for DFO |
createAB
![[top] [ top ]](/images/arr1red-up.gif) Description
Description
This tool creates association information and stores it in Association Blocks (ABs). It forms the central
part of the daily workflow. Find its place in the workflow here. Find a general description of the Association Block concepts here.
The tool uses ABbuilder as central association engine. That
tool uses the OCA framework, which is software developed by SEG for the purpose
of Organization, Classification
and Association of data. Other dfos tools making use of this framework are filterRaw and calChecker.
The tool works on a pool of raw hdr data defined
by <DATE> and expected to be found under $DFO_HDR_DIR/$DATE.
It has two fundamental modes:
- CALIB for calibration data,
- SCIENCE for science data.
createAB is executed on
a selected DATE. It also has an incremental mode working for an
incomplete date, used for the incremental processing scheme implemented with autoDaily. More about operational modes here.
Apart from ABs, createAB also creates a text file AB_list_<MODE>_<DATE> under $DFO_MON_DIR. This is the list of created ABs, sorted by the proper execution sequence. That list is fundamental for the daily workflow steps following createAB.
The tool has three TOOL_MODES:
- AUTO (no interaction required); this is the standard mode
- WARN (asks for interaction when a warning is found); this mode is useful
for command-line execution
- INTER (fully interactive).
The option -a can be used to override the configured value and enforce running in AUTO mode, which is useful if createAB itself is wrapped in automatic control tools like autoDaily or processPreImg.
Per configuration, the filter tool filterRaw can
be called (in mode FILTER) before ABs are created. All filtered files will be hidden before
AB creation. So they won't spoil your VCAL list, nor produce any AB. If required, they
can be inspected under ${HIDE_DIR}/<date>.
![[top] [ top ]](/images/arr1red-up.gif) Fits
files vs. headers. The entire
information needed for the creation of ABs is available in the headers. No need for fits files at that stage.
Fits
files vs. headers. The entire
information needed for the creation of ABs is available in the headers. No need for fits files at that stage.
![[top] [ top ]](/images/arr1red-up.gif) Incremental
mode. This
option is designed for the fast data transfer. It is triggered by the option -i and available
for mode=CALIB. If set, the tool will first look for already existing CALIB ABs for the
specified date. Under certain conditions, it will then move all raw header files listed
in these ABs (RAWFILE section) to a temporary directory, and will associate only the
remaining headers. The conditions are:
Incremental
mode. This
option is designed for the fast data transfer. It is triggered by the option -i and available
for mode=CALIB. If set, the tool will first look for already existing CALIB ABs for the
specified date. Under certain conditions, it will then move all raw header files listed
in these ABs (RAWFILE section) to a temporary directory, and will associate only the
remaining headers. The conditions are:
- TRY_AGAIN_INCOMPLETE=YES: headers listed in complete ABs are not associated
again
- TRY_AGAIN_FAILED=YES: headers listed in successfully executed ABs are
not associated again
In other words, usually only new headers, and headers from incomplete or
failed ABs, will be exposed to createAB, all others will not create again (and not process
either). If the above configuration keys are set to NO, only new headers will be used for
AB creation. Thereby AB creation and processing can be triggered once per hour, minimizing
processing delays and avoiding unnecessary multiple AB creation. Using this option in routine
operations requires a properly configured autoDaily (ENABLE_INCREM=YES).
One raw type can trigger several actions, i.e. two or more ABs can have the same set of raw files. New with version 3.9: If at least one of these ABs is failing then the raw file headers listed in these ABs are not moved. With the next call of createAB, these ABs are created again. Only if all ABs are successful then headers are moved and ABs are not created again. This takes care of special use cases where at least one AB depends on pipeline products from other reduction steps that might only become available later (because the calibrations are not yet measured on Paranal). As a side effect, it can result in several successful executions of the same AB until all ABs with the same set of raw files are successful.
No VCALs. With the option -N, the tool uses only certified and renamed
master calibrations (MCALs) for association, no virtual calibrations (VCALs). It is available
for mode=SCIENCE only. This is a stronger association constraint than the configurable
flag SCI_VIRT_MCAL which gives only a warning in case of VCALs being associated. It is
available on the command-line or by configuration of the dfoMonitor (CREATEAB_VCAL=YES).
![[top] [ top ]](/images/arr1red-up.gif) Completeness
checks
Completeness
checks
In mode SCIENCE, a completeness check is made between the raw files as extracted
from the science ABs, and the science raw files as extracted from data_products. In
case of a difference, it raises an alert (email and command-line).That difference would
have the consequence that the corresponding science ABs could not be harvested. It would
be caused by an OCA configuration inconsistency or incompleteness.
![[top] [ top ]](/images/arr1red-up.gif) Re-creation
of ABs. The tool offers support for the re-creation of ABs. This might
become of interest if
Re-creation
of ABs. The tool offers support for the re-creation of ABs. This might
become of interest if
- virtual calibration products required for an AB did not process well,
hence that AB failed but could be saved with another association (use case "failed")
- virtual calibration products required for an AB were
rejected, hence that AB is invalid and must be created again with another association (use
case "invalid")
- the association rules, or recipe parameters, must be modified for an AB,
hence that AB must be processed again (use case "OCA")
For AB re-creation the tool is called with option -r (recreate,
or reprocess). Before calling, the user has to prepare the environment. The exact
measures depend on the use case:
- clean up VCAL directory, to get rid of
associations to unwanted virtual products. This step is offered within certifyProducts if
an AB is rejected, but not if an AB failed: then the user must take care of deletion
(use cases failed, invalid).
- hide a bad raw file header, to avoid unwanted associations: if a certain
raw file is part of the cascade, it would create unwanted ABs and unwanted virtual calibrations
as soon as createAB is called again. Either you hide a header manually
(move it somewhere else and move it back later), or use filterRaw and hideFrames to
do it in the proper way (use cases failed,
invalid).
- clean up MCAL directory and NGAS: this is necessary if an unwanted real
product has already been certified, moved, renamed and archived. This might happen if
a quality issue is detected at some later point. These steps are not supported by dfos
tools, except for NGAS deletion with cdbDelete (use
case invalid).
- If you want to create and process all ABs for a given date, you
actually do not need the re-creation mode since it is designed for interactive AB selection (use
case OCA).
In re-creation mode the tool goes through the standard workflow
but has certain additional parts with dialogues marked by "[RECREATE]". These
are interactive by design. For better distinction, you may want to call the option '-a'
in addition, thereby forcing the tool to be interactive only in the re-creation parts.
You can do the call on the command line, or use the interactive call on the AB monitor
("recreate
ABs") which is more comfortable.
After having created all ABs for the specified date and mode, a window pops
up with the complete AB list and you are prompted for the AB selection. This is an editor
session, with your favorite editor as configured in config.createAB,
in the key
RECREATE_EDITOR. Mark your selection by a leading 'R<blank>', e.g.:
| R GIRAF.2010-05-05T23:08:41.611_tpl.ab |
NFLT |
Argus_L881.7 |
[selected] |
GIRAF.2010-05-05T23:16:14.696.ab
|
STD_ARG |
Argus_L881.7 |
[not selected] |
You do not need to care about hidden dependencies, just select the ABs you
know you want to reprocess. Next, the tool analyses dependencies. Dependent ABs are
those that are using products from a selected one. The tool displays all selections and
dependencies in a second interactive editor session:
| R
GIRAF.2010-05-05T23:08:41.611_tpl.ab |
NFLT |
Argus_L881.7 |
[selected] |
| \_GIRAF.2010-05-05T23:19:21.308.ab |
WAVE |
Argus_L881.7 |
[AB depending on the
selected one] |
At that stage you could also remove selections or dependencies, but removing
dependencies usually makes no sense.
After the prompt, the tool creates a new AB list called AB_list_<mode>_<date>_recreate.
As the last step, createAB then calls createJob (this step is different from
normal execution!) because this needs to be done with a special syntax: createJob
-m <mode> -d <date> -r <job_ID> where job_ID is <mode>_<date>_recreate, thereby
creating the usual job files execAB etc. with suffix _recreate,
containing only the selected ABs. This is a mechanism to protect any standard job file
which might exist and contain the complete list. These job files are written into the standard
JOBS_NIGHT, at which point the tool stops, leaving the execution of JOBS_NIGHT to the
user.
The jobs will execute as normal and will deliver the usual products which are then available
for review together with any other pre-existing products.
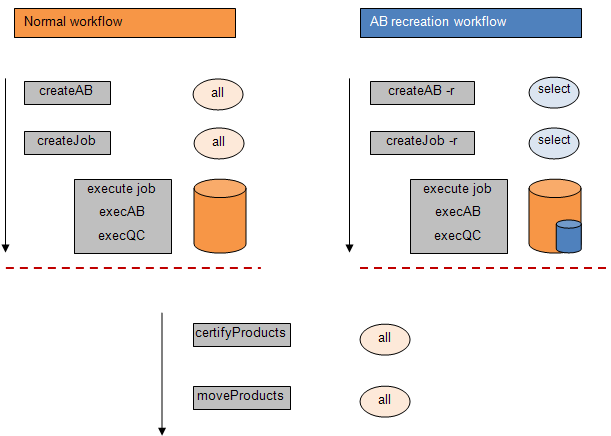 [click
to enlarge]
[click
to enlarge]
![[top] [ top ]](/images/arr1red-up.gif) Architecture. The tool is a shell-script wrapper calling the DFS
tool ABbuilder. createAB provides all functionality which is specific to DFO while ABbuilder is
the central engine largely independent of DFO-specific settings. Strictly speaking ABbuilder again
is a shell-script wrapper around a central java application:
Architecture. The tool is a shell-script wrapper calling the DFS
tool ABbuilder. createAB provides all functionality which is specific to DFO while ABbuilder is
the central engine largely independent of DFO-specific settings. Strictly speaking ABbuilder again
is a shell-script wrapper around a central java application:
![[top] [ top ]](/images/arr1red-up.gif) Association
logs. createAB displays the association log to the console, and also into
separate text files. These come per AB, have the same root name and the extension .alog,
and are found under $DFO_AB_DIR. In particular they contain all associated MCALIBs/MASSOCs,
their DELTA_T values, the configured DELTA_T values (validity), plus warnings: warnings
if the configured threshold value has been violated, warnings if a required mcalib could
not be found, and warnings if a SCIENCE AB has virtual calibrations associated (if keys
SCI_VIRT_MCAL | SCI_VIRT_MASS configured as YES). These entries are scanned by the AB
monitor tool getStatusAB and
get displayed.
Association
logs. createAB displays the association log to the console, and also into
separate text files. These come per AB, have the same root name and the extension .alog,
and are found under $DFO_AB_DIR. In particular they contain all associated MCALIBs/MASSOCs,
their DELTA_T values, the configured DELTA_T values (validity), plus warnings: warnings
if the configured threshold value has been violated, warnings if a required mcalib could
not be found, and warnings if a SCIENCE AB has virtual calibrations associated (if keys
SCI_VIRT_MCAL | SCI_VIRT_MASS configured as YES). These entries are scanned by the AB
monitor tool getStatusAB and
get displayed.
ATAB files (or tab files). These are text files
with one line in tabular form (hence their name). Their name is <AB_ROOT>.tab. They
contain the complete information about a particular AB as displayed on the AB monitor.
It has 33 entries, like e.g. processing status, display color, execution time, HC flag,
name of processing log, score results. Each of these ATAB files is generated by createAB,
and then updated throughout the workflow by the various tools (processAB, processQC, certifyProducts,
harvestAB). At any stage in the workflow, the tool getStatusAB can
read the tabular information and transform it directly into HTML code, without further
queries for information. This is a very fast way for displaying the actual AB monitor,
and very efficient since information is only updated when needed, not every time when the
tool getStatusAB is called.
ATAB files are created and maintained in $DFO_AB_DIR along with the ABs,
and then move to their final destination $DFO_LOG_DIR/$DATE. They are not exported
to qcweb since their only value is technical (speed up the AB monitor). They do not
contain any information beyond the ABs and format information for the AB monitor.
![[top] [ top ]](/images/arr1red-up.gif) Output
Output
The tool creates the following output:
- ABs under $DFO_AB_DIR (extension .ab)
- AB logs under $DFO_AB_DIR (extension .alog)
- a list of ABs, $DFO_MON_DIR/AB_list_<MODE>_<DATE>,
read by other tools supporting ABs
![[top] [ top ]](/images/arr1red-up.gif) How to use
How to use
Type createAB -h | -H | -v for on-line help about createAB / ABbuilder, and version.
Type
createAB -m CALIB -d 2016-12-30
to create ABs for mode CALIB and date 2016-12-30;
createAB -m SCIENCE -d 2016-10-30
-c test_config.createAB
to create ABs for mode SCIENCE and date 2006-10-30, with a non-standard
config file test_config.createAB (mcalib pool is defined by $N_MCAL_LIST and the latest date in $DFO_CAL_DIR, which usually is the one from -d or a few days later);
createAB -m SCIENCE -d 2006-10-30 -D 2006-11-02
to create ABs these ABs with a pool of mcalib files starting at date 2006-11-02
and going backwards by $N_MCAL_LIST days as configured;
createAB -m CALIB -d 2010-09-30 -i
to create CALIB ABs incrementally, i.e. only if they are did not exist before;
createAB -m CALIB -d 2010-09-30 -r [
-a ]
for re-creation of selected ABs.
Status
The tool writes the status values cal_AB or
sci_AB into DFO_STATUS. It also writes a timestamp into each created AB, and creates association log files under $DFO_AB_DIR which are scanned by getStatusAB and distributed together with the ABs.
![[top] [ top ]](/images/arr1red-up.gif) Configuration files
Configuration files
The
tool has a configuration which is somewhat more complex than for most other DFOS tools.
All relevant configuration is kept under $DFO_CONFIG_DIR/OCA/:
- config.createAB which has the createAB tool configuration
- OCA_macro.h: general macros
- <instr>_macro.h: instrument specific macros ( <instr> is short
for $DFO_INSTRUMENT)
- log4j.cf which is a java logging configuration file
- a set of config files in OCA syntax (these are NOT required for OPSHUB installations!):
- <instr>_classification.h : all classification configuration
- <instr>_organisation.h: all organization configuration
- <instr>_association.h: all association configuration
- <instr>.h: all include information (auto-created by the tool,
listed here for completeness
Find here more details about the OCA syntax.
The user may want to specify a non-standard config file by using the -c option
(the standard one being config.createAB), e.g. to handle pre-imaging association in addition
to standard DFO association. That file must also reside under $DFO_CONFIG_DIR/OCA.
The DFOS configuration file config.createAB has the following structure:
| KEY |
Description |
Example |
Comments |
| 1. Tool and general parameters |
| CONF_VERSION |
Version for the set of configuration files |
config.createAB_v2.0 |
|
| TOOL_MODE |
Execute mode (INTER or ERROR or AUTO) |
ERROR |
AUTO: automatic mode, no interruption; INTER: interactive;
ERROR: partly interactive (interaction only for missing calibrations);
Note: can be overridden by option -a[utomatic] at runtime |
| FILTER_RAW |
turn on or off filterRaw |
YES
| YES|NO (optional, default: NO). If Y, filterRaw is called before AB
creation, and matching files are removed |
| PGI_PREPROC |
use optional plugin within procedure launchAB |
<name> or NONE |
optional, default NONE |
| PGI_POSTPROC |
use optional plugin just before section 4 |
<name> or NONE |
optional, default NONE; DATE and MODE exported |
| PGI_FINAL |
use optional plugin just before the tool finishes |
<name> or NONE |
optional, default NONE; DATE and MODE exported |
| ACCEPT_060 |
accept science files with run_id 60./060./0060. |
YES | NO (default) |
if you exceptionally want to create ABs for these SCIENCE data, configure
as YES |
| XTERM_GEOM |
|
100x25+1000+1000 |
size and position of xterm with RECREATE_EDITOR call (default: 100x25+1000+1000
[bottom right]) |
| 1.2 General parameters |
| GEN_CALDIR |
$CAL_DIR directory for general (static) calib files |
$DFO_CAL_DIR/gen |
DFO convention |
| N_MCAL_LIST |
number of nights to be scanned for mcalib_list |
5 |
to be fine-tuned for your instrument |
| N_VCAL_LIST |
number of nights to be contained in vcalib_list |
5 |
to be fine-tuned for your instrument |
| CAL_N_MCAL_LIST |
like N_MCAL_LIST, but for
CALIB mode only (for SCIENCE, always N_MCAL_LIST applies) |
5 |
optional; default: N_MCAL_LIST |
| CAL_N_VCAL_LIST |
like N_VCAL_LIST, but for
CALIB mode only (for SCIENCE, always N_VCAL_LIST applies) |
5 |
optional; default: N_VCAL_LIST |
| DRS_TYPE |
Type of DRS |
CON | CPL | INT |
used to control recipe execution (CON: CONDOR; INT: internal parallelization with likwid-pin) |
| SCI_VIRT_MCAL |
give a warning in the alog (*NOK) if a SCIENCE AB has VIRTUAL mcalibs associated (in the MCALIB section) |
YES | NO |
YES makes sense if SCIENCE ABs are processed (since this AB is doomed to fail). Default is YES, key is optional. Key is evaluated for mode SCIENCE only. |
SCI_VIRT_MASS
|
same as SCI_VIRT_MCAL, for MASSOC section |
YES | NO |
same as SCI_VIRT_MCAL, for MASSOC data |
| SUPPRESS_VIRT |
list of PRO.CATGs for which no VIRTUAL alert is given for the science ABs |
SKY_LINES |
optional key (multiple lines supported) |
| PROD_STEP2 |
name of ACTION for step 2 ("child") ABs |
e.g. ACTION_FF_EXTSPECTRA |
optional key used in case you want to launch a second action (create a second kind of ABs) which has no triggering RAW_TYPE. Check here for details. Multiple lines are supported since v3.9. |
| TRY_AGAIN_INCOMPLETE |
for incremental mode: create ABs again if previous version was incomplete |
YES | NO |
default: YES |
| TRY_AGAIN_FAILED |
for incremental mode: create ABs again if previous version failed |
YES | NO |
default: YES |
| 2. Additional configuration |
| PACK_ADD |
Non-FITS products to be packed into ANCillary files:
PS = ps QC files
GIF = gif QC files
PNG = png files
JPG = jpg files
|
VALUE = PS: type of product, see above;
EXTENSION = ps.gz: identification is based on product root name plus EXTENSION
DIRECTORY = $DFO_PLT_DIR: directory (DFO convention) of product |
further values could be supported upon suggestion |
| NOCHECK |
YES | NOCHECK |
default: YES |
Used by updateAB v1.3.1 and higher; NOCHECK turns off checking for virtual/real status of calibrations in the ABs. This is useful for some special applications but not in usual dfos workflows. |
![[top] [ top ]](/images/arr1red-up.gif) Cascade configuration
Cascade configuration
The configuration of the cascade follows the matrix scheme which has from
left to right all raw types in their logical sequence, and from top to bottom the process
flow from raw to product, involving the grouping rule, the recipe, the required input mcalibs
and the predicted products. The classification defines the raw types and is configured in <instr>_classification.h. The grouping rules are configured in <instr>_organisation.h. The association rules are configured in <instr>_association.h.
There are currently the following cascade types supported by createAB:
- classical "simple" cascade: one raw_type, one action
- cascade with multiple actions: one raw_type, two actions
- cascade with parent-child actions: one raw_type triggers one action, second action is triggered by products of the first action (not by a raw_type)
Not supported types are e.g. cascades with raw data from different nights or different OBs.
Simple cascade: one action per raw_type
The classical cascade is configured by one entry block per raw_type, in each of the OCA config files. The sequence of raw_type definition is evaluated by the tool and should reflect the processing sequence (left to right ordering).
Cascade with multiple actions
An example for that type could be jittered science observations: create and process an AB per single raw file first, e.g. in order to derive QC information. Then create and process an AB per TPL set, in order to create the de-jittered product. Both actions are triggered by the same raw_type.
That cascade is configured in the same way as the simple cascade. In addition, the second action for a given raw_type is entered by additional blocks in the OCA config files.
Cascade with step1-step 2 ABs ("parent-child ABs")
This type has ABs which have no associated raw_type but virtual product files as input. A classical example is the treatment of imaging zeropoints:
| BIAS |
FLAT |
STD
(step1) |
(step2) |
mbias |
|
|
|
|
mflat |
|
|
|
|
single_zp |
|
|
|
|
night_zp (made from single_zp1, single_zp2) |
In step1, all raw input STD files create corresponding ABs (step1 or parent ABs). The products of those (here called 'single_zp', typically all per setup from the night) need to be combined in another (step2, child) AB which has no input raw files and produces a single night_zp product.
That case cannot be handled in the classical, raw-type driven way. It is configured in config.createAB using the PROD_STEP2 key.
- Define the RAW_TYPE for step1 (say: STD) in the usual way in the C part of the OCA rules; also define another RAW_TYPE (say NIGHT_ZP), using the appropriate PRO.CATG of the step1 product instead of DPR.TYPE;
- then define two ACTIONs in the O part (ACTION_STD and ACTION_NIGHT_ZP) with different extensions (e.g. TPL_A,tpl for the step1 action, and NIGHT,zp for the step2 action);
- then define both ACTIONs in the last part of the OCA files.
Note that the name of the step2 ACTION must be ACTION_<procatg> where procatg is the PRO.CATG triggering the second step. In config.createAB, configure PROD_STEP2 as ACTION_NIGHT_ZP.
If PROD_STEP2 is set, the tool calls ABbuilder twice:
- first, it calls ABbuilder with the purpose to create the parent (step1) ABs and from them the virtual calibration products (single_zp in the above example). These are then copied into $RAW_DIR, with extension .HDR. Thereby they become 'pseudo-raw files' which the tool then can handle in the standard way.
- second, it calls ABbuilder in the usual way. Now the input directory contains all input file information (raw and pseudo-raw) and can create both parent and child ABs.
The HDR files are deleted afterwards. This mechanism works both for hdr and fits input files. Since version 3.9, multiple definitions of PROD_STEP2 are supported.
![[top] [ top ]](/images/arr1red-up.gif) Workflow description
Workflow description
(This description applies to the DFOS and PHOENIX installations. For OPSHUB installations, the tool is only called internally and in a simplified way.)
1. Filter input data pool
- if FILTER_RAW=YES: call filterRaw -m FILTER -H, to remove the found files from further
processing. Corresponding headers are also moved, but moved back to their original folder at
the end.
2. Pre-compile all OCA configuration files into a final <instr>.RLS file (using gcc)
3. Prepare links under $DFO_CAL_DIR/MCAL to all mcalibs
as defined by $N_MCAL_LIST and the latest date in $DFO_CAL_DIR. If option -D <SCANDATE> has
been set, the set is defined by this start date instead. Supported are both fits and hdr
files in $DFO_CAL_DIR.
4. Only if PROD_STEP2 is set:
4.1 call ABbuilder to create virtual product
headers; call PGI_PREPROC if configured
4.2 copy the virtual product headers as .HDR files into RAW_DIR; no ABs are created at that step
5. Only with option -i (incremental) and mode=CALIB:
5.1 check for pre-existing ABs
5.2 move their headers to temporary space, unless INCOMPLETE (if TRY_AGAIN_INCOMPLETE=YES)
or FAILED (if TRY_AGAIN_FAILED=YES)
5.3 have only new headers in input data pool, plus the ones from step 5.2
6. Call ABbuilder:
6.1 Classify input data pool
- read fits keys as defined in <instr>_classification.h to classify each raw file into RAW_TYPE | DO_CLASS. (Note: PACK_DIR is obsolete and can be removed.)
6.2 Organize input data into groups
- get raw match keys from <instr>_organisation.h
- get grouping rules (e.g. SINGLE, TPL_A, TPL_D etc.)
6.3 Find association
per group, reading rules in <instr>_association.h
- evaluate MCALIB file match keys to find all calibration
products required for processing
-
evaluate MASSOC match keys to find additional mcalib files (if any) useful for packing
-
evaluate RASSOC match keys to find
associated raw files (if any) useful for packing
- find recipe, recipe parameter
- find WAITFORs for CONDOR (from associated virtual calibrations)
-
set the AB status to 'created'
6.4 Create the ABs in a work directory ($DFO_AB_DIR/TMP)
- call PGI_PREPROC if configured
6.5 Add virtual calibrations as predicted from the ABs to $DFO_CAL_DIR/VCAL
7A. (normal operations, including incremental mode):
Verify
the ABs (ordered in a cascade as configured in <instr>_classification.h)
- scan for COMPLETE/INCOMPLETE flag
- create and display the association log (includes all found MCALIB/MASSOC with their DELTA_T values, and all missing ones)
- create the .tab files for speedy AB monitor page
- edit the RAW_MATCHKEY section to suppress UNDEFINED entries (typical
of mutually exclusive keys e.g. in VIMOS or FORS2)
- for SCIENCE only: check if MCALIBs and MASSOCs have VIRTUAL calibrations (flag as
*NOK)
- if configured (TOOL_MODE=INTER/WARN), ask the user what to do in case of incomplete
ABs
- construct the list of all created ABs
- move ABs and AB logs to $DFO_AB_DIR
7B. (RECREATE=YES): same as 7A, plus
- offer the created ABs and
let the user decide which ones to select
- create the list of ABs on the selected ones and the ones depending on them
- move only those to $DFO_AB_DIR, the other ones are deleted
8A. (normal operations) After AB creation:
- if step 5 was applied (incremental mode), move all headers back to $DFO_HDR_DIR/$DATE
- apply PGI_POSTPROC if configured
- create directories in $DFS_PRODUCT
- manage the set of virtual calibrations (headers in $DFO_CAL_DIR/VCAL): remove outdated ones (defined by $N_VCAL_LIST)
- move back hidden headers to $DFO_HDR_DIR/$DATE
8B. (RECREATE=YES): same as 8A, plus
- call createJob and create the job files for the selected set of ABs (the
job files come with suffix _recreate)
9. call PGI_FINAL if configured
![[top] [ top ]](/images/arr1red-up.gif) Operational hints
Operational hints
- The incremental mode is the standard mode for calibrations of all instruments
- It complies with
other parallel activities like e.g. scheduled jobs in JOBS_NIGHT (inc. processing other
ABs, QC reporting etc.). This just increases the load on the machine but does
not break anything.
- In MODE=SCIENCE, all files with OBS_PROG_ID starting with 60./060./0060.
are filtered (not accepted) (unless ACCEPT_060=YES).
- You can use CAL_N_MCAL_LIST to have different calibration memory depths
for CALIB and for SCIENCE (which may make sense but is normally not required)
- You can call three plugins, one (PGI_PREPROC) directly after calling ABbuilder
(works in the temporary AB directory, used e.g. to edit setup keys), one (PGI_POSTPROC)
on the final ABs (after editing them and moving them to $DFO_AB_DIR), one (PGI_FINAL) just
at the end. All plugins are optional. If defined, they must exist in $DFO_BIN_DIR.
| Last update: April 26, 2021 by rhanusch
|
